8.15 Look for Gaps in Coverage as a Sign of Missed Exons
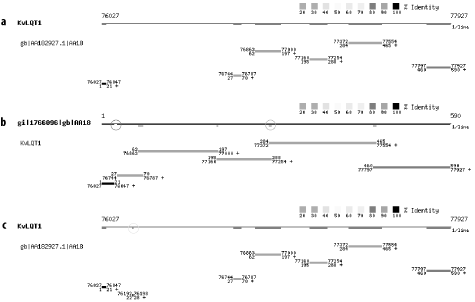
The seeding parameters and alignment thresholds may prevent short or highly divergent exons from appearing in BLAST reports. Figure 8-8a shows an alignment between a genomic query and an EST. Most alignments overlap by a few bp, except for the 2 at the 5´ end (left side). Gaps and overlaps in coverage are easier to see by using the reciprocal search shown in Figure 8-8b. To find the missing 7-bp exon in Figure 8-8c, use bl2seq (see Chapter 13) with the following command line:
bl2seq -i est -I 21,29 -j genomic -J 76047,76744 -pblastn -W 7
The -I and -J parameters let you select a specific region of each sequence. What you've done is a BLASTN search between the missing part of the EST and the region between the alignments.
Figure 8-8. Finding missed exons: (a) an alignment between a genomic query and EST, (b) the reciprocal alignment showing a gap (d) and overlap (e) in coverage, (c) the tiny missed exon can be found (f) by changing the word size to 7